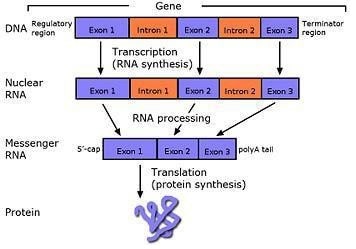
The following are 3 DNA fragments that together contain one gene. Follow the steps below to convert the gene code from DNA into a protein.
Fragment A: AATCTACCGAGGCCGTCCGCCGCCATAT
Fragment B: TTTGCCCCGGGATTTCCCATCCCAATTGGGG
Fragment C: CGCCCCTCAGAAGTTGGTTCAGACGTTGGC
Procedure:
- Find the correct order the 3 fragments must be in. Recopy the DNA code for the GENE in order from start to stop. ✓
- Transcribe the GENE into mRNA. ✓
- Remove the 2 introns which include bases 10-15 and 25-30. ✓
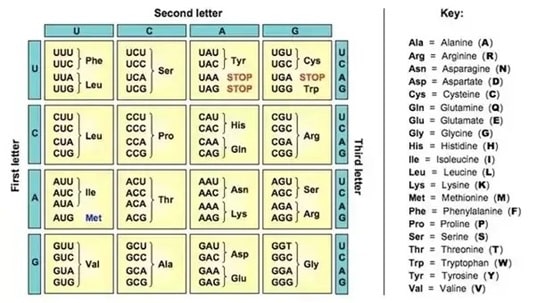
- Separate the codons of the mRNA and Translate into an amino acid sequence. ✓ Results: Show all work and use references.
Analysis:
Which fragments were the beginning, middle, and end? Justify.
At what point in the process are introns removed? What is the name of the enzyme responsible for this excision?
How many amino acids are in the final protein?
Contrast (at least 3 factors) protein synthesis between prokaryotes and eukaryotes. If you used the Edman method (p.48) on this protein to find the amino acid sequence and worked backward through mRNA to DNA, would you find the exact DNA sequence as above? Explain why or why not?
What are operons? Explain with an example.
Procedure/Results: Step 0.

Analysis:
- Which fragments were the beginning, middle and end? Justify.
Fragment A was the beginning as it contains the starting codon AUG. This signifies the starting of the code. The middle fragment was fragment C as fragment C had no starting or ending codons to signify a specific placement of the fragment. Placing fragment C after fragment A and before fragment B allowed for the end code UAG to appear in the code. Placing fragment B at the end was due to the original appearance of the end codon UAG and allows for a seemingly clear end to the code as well as removing any extra codons that follow the initial end codon.
- At what point in the process are introns removed? What is the name of the enzyme responsible for this excision?
Introns are removed in the pre-mRNA splicing process and are exported to the cytoplasm whilst exons are then to be joined to form a coding sequence. Which transforms this RNA into “mature” mRNA is then ready for translation. The RNA-splicing enzyme is known as endonuclease. It is an evolutionarily conserved enzyme responsible for the excision of introns from nuclear transfer RNA and all archeal RNAs.
- How many amino acids are in the final protein?
There are a total of 20 amino acids in the final protein including the starting amino acid. The 20 amino acids consist of the starting amino acid Methionine, and various quantities of Alaine, Proline, Glycine, Valine, Phenylalanine, Asparagine, Arginine, Lysine, Leucine and more. There is in all technicalities only a total of 20 amino acids within the final protein as stop codons are not amino acids and if the stop codons were to be included as amino acids, then the sequence would considerably have 21 amino acids rather than 20 (W2).
Contrast (at least 3 factors) protein synthesis between prokaryotes and eukaryotes. Transcription and translation happen at the same time in prokaryotes, but can happen at different times in eukaryotes. This is because prokaryotes do not have membrane bound organelles whereas eukaryotes do. Eukaryotes have transcription occur in the nucleus and translation occurs in the cytoplasm of the cell, and prokaryotes have translation alongside transcription occur in the cytoplasm (W4). Eukaryotes do not have operons unlike prokaryotes as in eukaryotes, gene regulation can be complex so it becomes hard to fit genes for regulation (W1).
Operons in prokaryotes can essentially allow the concerted expression of function related genes as simpler polycistronic transcripts. This is rare in eukaryotes as each gene usually drives expression of its own independent messenger RNAs (W5). In Eukaryotes, most mRNA undergoes splicing before it can be translated, so eukaryotic genes can require mRNA that are spliced, as eukaryotic genes can contain introns that are removed prior to protein synthesis. Prokaryotes don’t perform post transcriptional modification, as they do transcription and translation at the same time and mainly has to do with timing.
- If you used the Edman method (p.48) on this protein to find the amino acid sequence and worked backwards through mRNA to DNA, would you find the exact DNA sequence as above? Explain why or why not?
Unfortunately no, the Edman method would not give you the same DNA sequence as above. This is due to the multiple codons that code for the same amino acid, for reference GUU, GUC, GUA, and GUG, are all code for the amino acid Valine. Now, because of this, the Edman method could not simply distinguish which one of the codes was present in the initial mRNA sequence (W6). Resulting, when transcribing from mRNA back to DNA, it would be simply impossible to know with certainty which codons were used to create the amino acid. Additionally, a large contrast with the Edman method is the presence of introns. The introns that are removed after the creation of the mRNA strand would not be included in the Edman method and would then also not be included in the recreation of the initial DNA sequence.
- What are operons? Explain with an example.
An operon is a group or cluster of genes that are transcribed together to give a single messenger RNA molecule that encodes multiple proteins. There are multiple components of an operon, including the operator and it’s interactions with a repressor. Usually, the repressor will bind to the operator to inhibit transcription of the genes encoded in the operon until specific criteria is met. They are found in prokaryotic cells and can affect the expression of operons. Operons can all be controlled by the same regulatory system and are typically expressed together. For an example of an operon, a lac operon is a great example of some of the possibilities they can hold. For a lac operon, it contains three different genes, lacZ, lacY, and lacA, and are controlled under one operator and transcribed as a single mRNA. The repressor will bind to the operator when lactose is absent from the cell. This will prevent RNA polymerase from transcribing the genes located on the lac operon. However, when lactose is present, it acts as an inducer and binds to the lac repressor. This binding causes the repressor to turn off, causing it to no longer bind to the operator region. This allows RNA polymerase to bind uninterrupted to the promoter region, allowing the genes located on the lac operon to be transcribed (B1). These genes within the lac operon allow for the prokaryotic cells to synthesize lactase when in the presence of lactose, which is can use as a food source to be used in glycolysis (W3).
References:
(W1) Byju’s. (2023). Is operon present in eukaryotes? Byju’s. Retrieved April 18, 2023, from
https://byjus.com/question-answer/is-operon-present-in-eukaryotes/
(W2) Cheriyedath, S. (2019, February 26). START and STOP Codons. News Medical. Retrieved April 18, 2023, from h ttps://www.news-medical.net/life-sciences/START-and-STOP-Codons.aspx (W3) Khan Academy. (2017). The lac operon (article). Khan Academy. Retrieved April 18, 2023, from
| (W4) ibreTexts. (2022, December 24). 7.6C: Prokaryotic Transcription and Translation Are Coupled. Biology LibreTexts. Retrieved April 18, 2023, from https://bio.libretexts.org/Bookshelves/Microbiology/Microbiology_(Boundless)/07%3A_Microbial_Genetics/7.06%3A_Translation-_Protein_Synthesis/7.6C%3A_Prokaryotic_Transcription_and_ Translation_Are_Coupled |
https://www.khanacademy.org/science/ap-biology/gene-expression-and-regulation/regulation-of- gene-expression-and-cell-specialization/a/the-lac-operon
(W5) National Library Of Medicine. (2019, February 21). Eukaryotic Acquisition of a Bacterial Operon – PMC. NCBI. Retrieved April 18, 2023, from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7295392/
(W6) PSIBERG. (2022, September 17). Edman Degradation: Chemistry, Steps, Limitations, Uses. PSIBERG. Retrieved April 18, 2023, from https://psiberg.com/edman-degradation/ (B1) Di Giuseppe, M., & Fraser, D. (2002). Nelson Biology 12. Nelson Education Limited.